Restriction enzymes are used in biotechnology to cut DNA into smaller strands in order to study fragment length differences among individuals. This is referred to as restriction fragment length polymorphism (RFLP). They're also used for gene cloning. Knowledge of these unique areas is the basis for DNA fingerprinting..
Hereof, how are restriction enzymes used in genetic engineering?
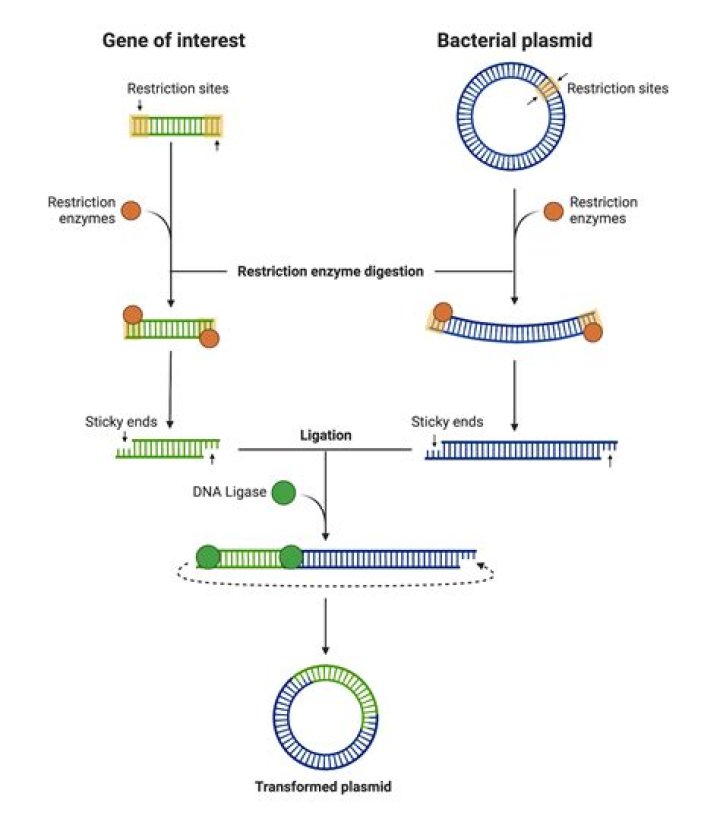
Restriction Enzymes are enzymes (naturally found in bacteria) that cut DNA at specific DNA sequences known as recognition site. Restriction Enzymes are useful in genetic engineering as they can be used to cut plasmids to produce 'sticky ends' (ends that are cut through a zig zag line, as shown in the above picture).
Likewise, what is the role of restriction endonuclease in biotechnology? Restriction endonucleases are enzymes that cut double-stranded DNA at very specific recognition sites. They were originally discovered in bacteria that use them to restrict the growth of viruses but are now among the workhorse enzymes of biotechnology and recombinant DNA research.
Considering this, how are restriction enzymes and ligase used in biotechnology?
Restriction enzymes are DNA-cutting enzymes. DNA ligase is a DNA-joining enzyme. If two pieces of DNA have matching ends, ligase can link them to form a single, unbroken molecule of DNA. In DNA cloning, restriction enzymes and DNA ligase are used to insert genes and other pieces of DNA into plasmids.
What are restriction enzymes used in?
Restriction enzymes. In the laboratory, restriction enzymes (or restriction endonucleases) are used to cut DNA into smaller fragments. The cuts are always made at specific nucleotide sequences. Different restriction enzymes recognise and cut different DNA sequences.
Related Question Answers
What does HindIII stand for?
HindIII (pronounced "Hin D Three") is a type II site-specific deoxyribonuclease restriction enzyme isolated from Haemophilus influenzae that cleaves the DNA palindromic sequence AAGCTT in the presence of the cofactor Mg2+ via hydrolysis.Which enzyme is useful in genetic engineering?
DNA ligases also play active part in processes such as DNA replication and recombination. These enzymes are widely used in genetic engineering for the production of hybrid DNA. Since ligase enzymes join DNA fragments or seal the nicks in the chain, they are called molecular structures.How do you choose restriction enzymes?
When selecting restriction enzymes, you want to choose enzymes that: - Flank your insert, but do not cut within your insert.
- Are in the desired location in your recipient plasmid (usually in the Multiple Cloning Site (MCS)), but do not cut elsewhere on the plasmid.
What is the source of restriction enzymes?
Bacterial species are the major source of commercial restriction enzymes. These enzymes serve to defend the bacterial cells from invasion by foreign DNA, such as nucleic acid sequences used by viruses to replicate themselves inside a host cell.How are enzymes named?
Enzymes are named by adding the suffix -ase to the name of the substrate that they modify (i.e., urease and tyrosinase), or the type of reaction they catalyze (dehydrogenase, decarboxylase). Structurally, the vast majority of enzymes are proteins. Also RNA molecules have catalytic activity (ribozymes).How are restriction enzymes useful to humans?
The ability of restriction enzymes to reproducibly cut DNA at specific sequences has led to the widespread use of these tools in many molecular genetics techniques. Restriction enzymes can be used to map DNA fragments or genomes.Why is the same restriction enzyme used?
The principle is simply that, if two different DNA molecules are cut with the same restriction enzyme, both will produce fragments with the same complementary sticky ends, making it possible for DNA chimeras to form.How do bacteria protect themselves from restriction enzymes?
The restriction enzymes in bacteria function to defend themselves against invading viruses (bacteriophages). Bacteria prevent eating away their own DNA by masking the restriction sites with methyl groups ( CH3 ). Methylation of DNA is a common way to modify DNA function and bacterial DNA is highly methylated.What is a ligase reaction?
In molecular biology, ligation refers to the joining of two DNA fragments through the formation of a phosphodiester bond. An enzyme known as a ligase catalyzes the ligation reaction. In the cell, ligases repair single and double strand breaks that occur during DNA replication.What is the job of ligase?
DNA ligase is an enzyme that repairs irregularities or breaks in the backbone of double-stranded DNA molecules. It has important role in the process of DNA replication and DNA repair.Which is the first discovered restriction endonuclease?
In 1970, Hamilton O. Smith, Thomas Kelly and Kent Wilcox isolated and characterized the first type II restriction enzyme, HindII, from the bacterium Haemophilus influenzae.How does DNA ligase work?
DNA ligase is an enzyme which can connect two strands of DNA together by forming a bond between the phosphate group of one strand and the deoxyribose group on another. It is used in cells to join together the Okazaki fragments which are formed on the lagging strand during DNA replication.Which restriction enzyme produce blunt ends?
Eco RV is type II restriction endonuclease isolated from Escherichia coli which produces blunt ends by making a cut in the center of the nucleotide sequence GAT/ATC.Is the double stranded DNA in the form of fragments or is it intact?
The answer is YES. Restriction enzyme cuts a DNA double helix in smaller fragments but the double helical structure of fragments remain intact.What determines how DNA will be cut by a restriction enzyme?
Recognition of different nucleotide sequences determines how DNA will be cut by a restriction enzyme. Restriction sites are the sequences of cut nucleotides, but restriction maps are maps of the restriction sites.Why is ligase needed to make recombinant DNA?
Why is ligase needed to make recombinant DNA? Answer: Ligase is an essential enzyme within all cells that seals breaks in the phosphate-sugar backbone of DNA. During DNA replication it joins Okazaki fragments to create a continuous strand, and in cloning, it is used to join the various DNA fragments with the vector.Who discovered restriction enzymes?
Restriction enzymes were discovered and characterized in the late 1960s and early 1970s by molecular biologists Werner Arber, Hamilton O. Smith, and Daniel Nathans.How did EcoRI get its name?
EcoRI. EcoRI (pronounced "eco R one") is a restriction endonuclease enzyme isolated from species E. coli. The Eco part of the enzyme's name originates from the species from which it was isolated, while the R represents the particular strain, in this case RY13.Do humans have restriction enzymes?
The HsaI restriction enzyme from the embryos of human, Homo sapiens, has been isolated with both the tissue extract and nuclear extract. It proves to be an unusual enzyme, clearly related functionally to Type II endonuclease.